Flat JSON API Implementation
Summary
Both CCDA and FHIR formats present data in complex hierarchies with multiple levels of nesting. While Particle supports both of them, many of our customers wanted to simplify their initial implementation, or in some cases needed only specific subset of the data. To enable users, we created Flat JSON format. To learn more about the schema, please see Flat Output Format & Supported Data Sets
The Particle Flat JSON API provides patient medical record data in a simplified format. The endpoint groups patient data by human-concept datasets - e.g. labs, vital signs, encounters, medications, etc.
This documentation will help you get started with initiating Flat queries, polling for the query status, pulling the returned data, and leveraging the data in your clinical workflow.
Particle Flat JSON API Flow
The Particle Flat API is centered around the patient. Each query will begin with submitting a set of patient demographics, which we will register and return a unique ID for. Subsequently, you can use this Query ID to check the status of the query for completion, and once ready, pull patient data in Flat output format.
Below is a high-level view of the full workflow:
Webhook Flow
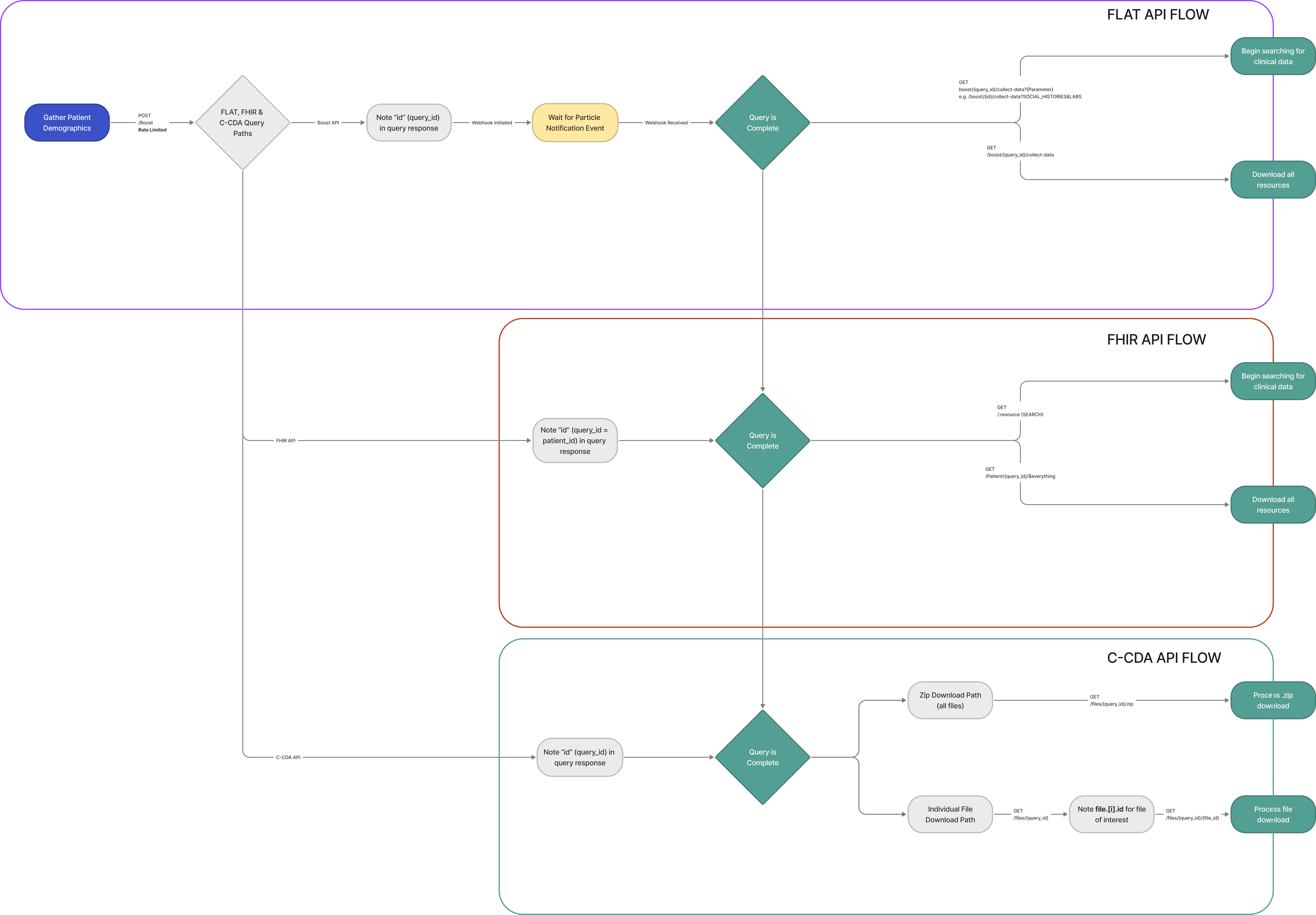
Polling Flow
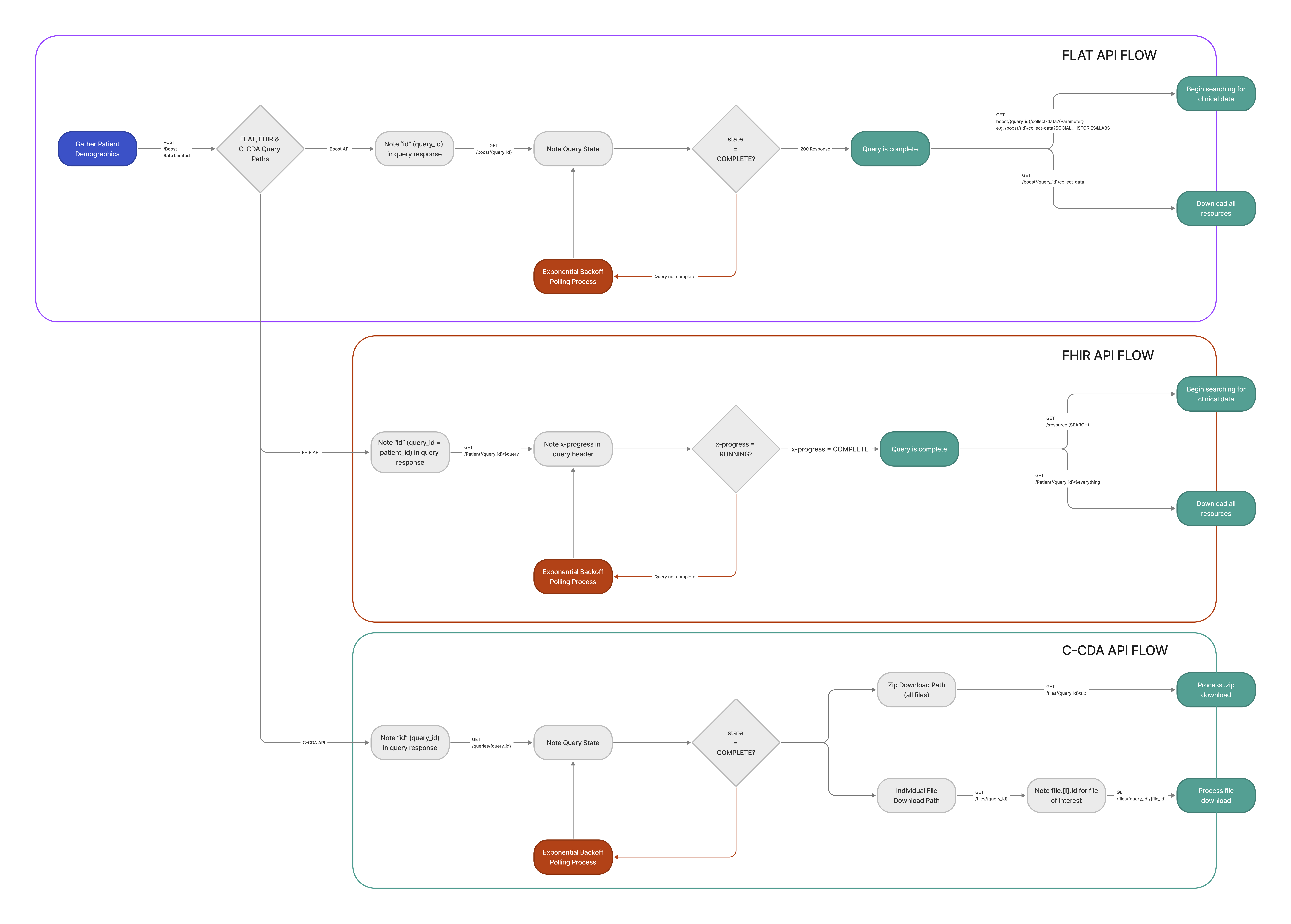
The following steps provide technical detail in how to perform the full query flow:
Initiating a Flat Patient Query
Submit Patient Demographics
The first step is to submit the patient demographics to the Flat endpoint to trigger a network query via the POST /flat route.
If this response is successful, you will receive a status code 200, as well as a unique Query_ID to use in subsequent steps in place of {id}.
curl --request POST \
--url 'https://api.particlehealth.com/flat' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json' \
--header 'content-type: application/json' \
--data '
{
"hints": [
"{additional_postal_codes}"
],
"address_city": "{city}",
"address_state": "{state}",
"date_of_birth": "{yyyy-mm-dd}",
"family_name": "{last_name}",
"gender": "{gender}",
"given_name": "{first_name}",
"postal_code": "{main_postal_code}",
"purpose_of_use": "TREATMENT"
}'
Example unique Query_ID Output: f34258bb-2b82-4b2e-940e-773ac823e387
The Flat Query_ID can also be used as the C-CDA Query_ID to pull C-CDA records or the Patient_ID for FHIR records
Retrieving the Flat Patient Query Status
The next step is to check the query status to determine if a query is complete and ready for data retrieval. You can opt to use our webhooks feature to be notified of query completion, or alternatively you can use the GET /flat/{id} route and poll this endpoint for a 200 status.
curl --request GET \
--url 'https://api.particlehealth.com/flat/{id}' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json'
Once you receive a 200 response from the GET /flat/{id} route, the query is ready for data retrieval. If you receive a 202 status code, this means the query is still processing.
The posted patient demographics will also be returned with a successful request to the GET /flat/{id} route to display which demographics an ID is associated with and to confirm that a set of demographics has been successfully POSTed.
Retrieving Flat Patient Query Data
Once the query is complete, the final step of the Flat process is to retrieve the data via theGET /flat/{id}/collect-data[parameters] route. You can specify which Particle Flat datasets to receive by specifying them in the [parameters].
Below is an example of how to retrieve the labs data set for a given Query ID:
curl --request GET \
--url 'https://api.particlehealth.com/flat/{id}/collect-data?LABS' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json'
The following section details the available Particle Flat datasets and how to format the parameters to request a single dataset, multiple datasets, or all datasets.
Selecting the Data Models Using the Parameters
To access any of the above datasets via the GET Flat data endpoint, include the dataset name as a parameter after the collect-data endpoint in the [parameters]:
curl --request GET \
--url 'https://api.particlehealth.com/flat/{id}/collect-data?LABS' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json'
For two word datasets, please concatenate the two word dataset name with an underscore in the [parameters]:
curl --request GET \
--url 'https://api.particlehealth.com/flat/{id}/collect-data?SOCIAL_HISTORIES' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json'
You can specify multiple datasets by concatenating the with an ‘&’ in the [parameters]:
curl --request GET \
--url 'https://api.particlehealth.com/flat/{id}/collect-data?SOCIAL_HISTORIES&LABS' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json'
Not specifying a dataset in the [parameters] will return all datasets available:
curl --request GET \
--url 'https://api.particlehealth.com/flat/{id}/collect-data' \
--header 'Authorization: {authorization_token}' \
--header 'accept: application/json'
This page contains further information on how to link the data models together to leverage the data for your clinical workflow.
Updated 8 months ago
